Welcome to the DepMap Portal
The goal of the Dependency Map (DepMap) portal is to empower the research community to make discoveries related to cancer vulnerabilities by providing open access to key cancer dependencies, analytical, and visualization tools.
Learn more about DepMap Learn more about PedDepDepMap is part of a public-private partnership called the DepMap Consortium. Learn more about becoming a member of the DepMap Consortium.
Explore the Portal
Updates and Announcements
26Q1 - Portal update
The 26Q1 release is now available! Please read the release notes for more details on new data and pipeline updates.
Release Highlights
- 25 new genome-wide CRISPR screens and Omics for adult and pediatric models
- Subtype tree updates for CNS models (October 9, 2025 OncoTree update)
- Updates to CRISPR pipeline for library correction
- Updates to Mutation pipeline for variant filtering and hotspots
- Context Manager update to allow for all metadata types to be used when defining contexts
- New Tool! GeneTEA uses natural language processing to uncover biology across the portal
Olink Proteomics dataset
We have profiled 161 cell lines covering 24 different cell line lineages using the Olink Explore HT platform. To learn more about Olink, please visit their website here.
Sanger Proteomics dataset
We’ve included a matrix of relative protein quantities from Gonçalves et al. 2022. The original data was downloaded from Cell Model Passports.
Read the full Release Notes
Explore More Ways To Use DepMap
Discover genetic and pharmacological dependencies
Explore MSI cancer dependency on WRN in Data ExplorerSee the Gene Dependency Overview page for Sox10
Prioritize tumor contexts and predictive biomarkers
Compare TRILACICLIB (CDK inhibitor) and RB1 expression in Data Explorer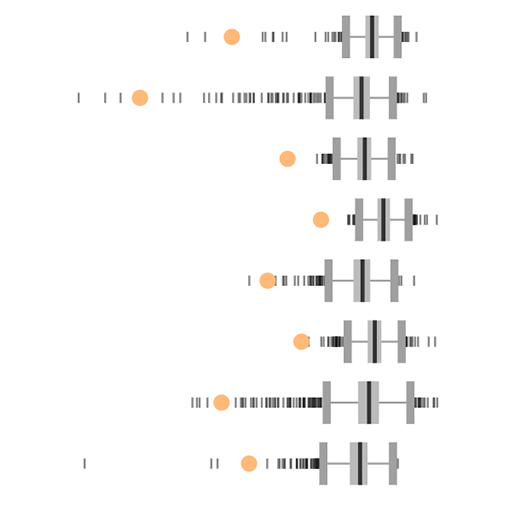
Browse our collection of over 2000 cancer models
Dive deeper into MCF7 on our Gene Dependency page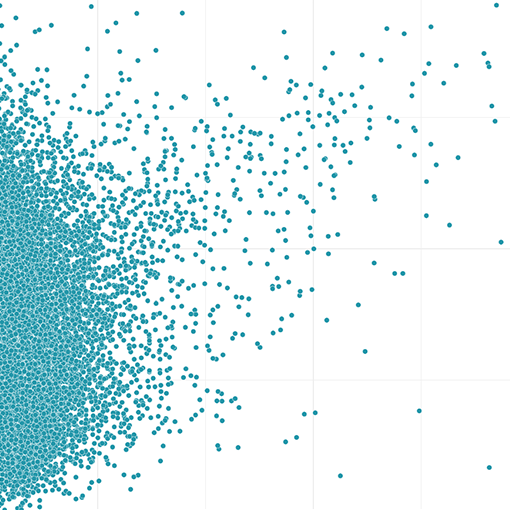
Discover interesting targets
Find the most selective and/or predictable gene dependencies in Target discoveryContribute Cancer Models and Datasets to DepMap