View Achilles data by searching for gene, cell line or lineage on the DepMap portal
Contact achilles@broadinstitute.org for more information
Project Achilles
Project Achilles is creating a catalog of essential genes
Achilles systematically identifies and catalogs gene essentiality across hundreds of genomically characterized cancer cell lines. We use genome-scale RNAi and, more recently, CRISPR-Cas9 genetic perturbation reagents to silence or knockout individual genes and identify those genes that affect cell survival. By linking these dependencies to the genetic or molecular features of the tumors, this project is providing the foundation for the "Cancer Dependency Map”.Highly standardized pooled genome-scale LOF screens
We use lentiviral-based pooled RNAi or CRISPR/Cas9 libraries in genome-scaled pooled loss-of-function (LOF) screening. This allows for the stable suppression of each gene individually in a subset of cells within a pooled format allowing for a cost-effective genome scale interrogation of gene essentiality. We developed a highly standardized parallel screening workflow able to handle a large number of screens and ensure that the data across cell lines are comparable. Rigorous quality controls, including multiple cell line fingerprinting steps and monitoring of cell/reagent representation throughout the screening process are performed to ensure the quality of the data.Computational Modeling for more accurate determination of gene essentiality
We have developed methods (DEMETER for RNAi screening, CERES for CRISPR screening) to computationally infer and subtract seed effects that arise for each shRNA. The resulting dataset is highly reliable and ready to be explored.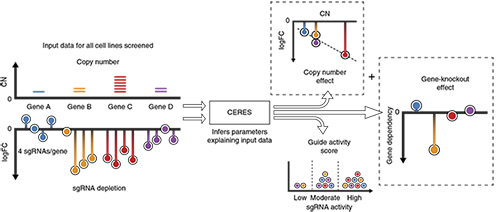
Schematic of the CERES computational model. CERES models the depletion values as a sum of gene-knockout and copy number effects, multiplied by a guide activity score parameter. CERES then outputs the values of the parameters that produce the highest likelihood of the observed data under the model.
Quarterly releases of data
Project Achilles is an ongoing effort aimed at screening more than 2000 cell lines of a variety of lineages in the next 5 years, including cell lines derived from both solid and hematopoietic tumors of pediatric and adult lineages. We are committed to making the Achilles data a resource for the scientific community. For unrestricted use of the latest datasets available please visit
our data page. Additional data is being generated and will be released on a quarterly basis pre-publication.
Legacy Datasets
As we update our computational methods to process the data, we will replace the older datasets with the latest versions. However, previously published but not currently used datasets are still stored on this portal.Select Publications
Aviad Tsherniak, Francisca Vazquez, Phillip G. Montgomery, Barbara A. Weir, Gregory Kryukov, Glenn S. Cowley, Stanley Gill, William F. Harrington, Sasha Pantel, John M. Krill-Burger, Robin M. Meyers, Levi Ali, Amy Goodale, Yenarae Lee, Guozhi Jiang, Jessica Hsiao, William F. J. Gerath, Sara Howell, Erin Merkel, Mahmoud Ghandi, Levi A. Garraway, David E. Root, Todd R. Golub, Jesse S. Boehm, & William C. Hahn. Defining a Cancer Dependency Map. Cell July 27, 2017. DOI: j.cell.2017.06.010
Andrew J. Aguirre, Robin M. Meyers, Barbara A. Weir, Francisca Vazquez, Cheng-Zhong Zhang, Uri Ben-David, April Cook, Gavin Ha, William F. Harrington, Mihir B. Doshi, Maria Kost-Alimova, Stanley Gill, Han Xu, Levi D. Ali, Guozhi Jiang, Sasha Pantel, Yenarae Lee, Amy Goodale, Andrew D. Cherniack, Coyin Oh, Gregory Kryukov, Glenn S. Cowley, Levi A. Garraway, Kimberly Stegmaier, Charles W. Roberts, Todd R. Golub, Matthew Meyerson, David E. Root, Aviad Tsherniak, & William C. Hahn. Genomic Copy Number Dictates a Gene-Independent Cell Response to CRISPR/Cas9 Targeting. Cancer Discovery 6, 914-929. June 3, 2016.
Glenn S. Cowley, Barbara A. Weir, Francisca Vazquez, Pablo Tamayo, Justine A. Scott, Scott Rusin, Alexandra East-Seletsky, Levi D. Ali, William F.J. Gerath, Sarah E. Pantel, Patrick H. Lizotte, Guozhi Jiang, Jessica Hsiao, Aviad Tsherniak, Elizabeth Dwinell, Simon Aoyama, Michael Okamoto, William Harrington, Ellen Gelfand, Thomas M. Green, Mark J. Tomko, Shuba Gopal, Terence C. Wong, Hubo Li, Sara Howell, Nicolas Stransky, Ted Liefeld, Dongkeun Jang, Jonathan Bistline, Barbara Hill Meyers, Scott A. Armstrong, Ken C. Anderson, Kimberly Stegmaier, Michael Reich, David Pellman, Jesse S. Boehm, Jill P. Mesirov, Todd R. Golub, David E. Root, & William C. Hahn. Parallel genome-scale loss of function screens in 216 cancer cell lines for the identification of context-specific genetic dependencies. Nature Scientific Data 1, Article number: 140035. September 30, 2014.
Mehmet Gönen, Barbara A. Weir, Glenn S. Cowley, Francisca Vazquez, Yuanfang Guan, Alok Jaiswal, Masayuki Karasuyama, Vladislav Uzunangelov, Tao Wang, Aviad Tsherniak, Sara Howell, Daniel Marbach, Bruce Hoff, Thea C. Norman, Antti Airola, Adrian Bivol, Kerstin Bunte, Daniel Carlin,2 Sahil Chopra, Alden Deran, Kyle Ellrott, Peddinti Gopalacharyulu, Kiley Graim, Samuel Kaski, Suleiman A. Khan, Yulia Newton, Sam Ng, Tapio Pahikkala, Evan Paull, Artem Sokolov, Hao Tang,1 Jing Tang, Krister Wennerberg, Yang Xie, Xiaowei Zhan, Fan Zhu, Broad-DREAM Community, Tero Aittokallio, Hiroshi Mamitsuka, Joshua M. Stuart, Jesse S. Boehm, David E. Root, Guanghua Xiao, Gustavo Stolovitzky, William C. Hahn, & Adam A. Margolin. A Community Challenge for Inferring Genetic Predictors of Gene Essentialities through Analysis of a Functional Screen of Cancer Cell Lines. Cell Syst. 2017 Nov 22;5(5):485-497.e3. doi: 10.1016/j.cels.2017.09.004. Epub 2017 Oct 4.
Xiaoyang Zhang, Peter S. Choi, Joshua M. Francis, Galen F. Gao, Joshua D. Campbell, Aruna Ramachandran, Yoichiro Mitsuishi, Gavin Ha, Juliann Shih, Francisca Vazquez, Aviad Tsherniak, Alison M. Taylor, Jin Zhou, Zhong Wu, Ashton C. Berger, Marios Giannakis, William C. Hahn, Andrew D. Cherniack, & Matthew Meyerson. Somatic super-enhancer duplications and hotspot mutations lead to oncogenic activation of the KLF5 transcription factor. Cancer Discov September 29 2017 DOI: 10.1158/2159-8290.CD-17-0532
Hubo Li, Brenton G. Mar, Huadi Zhang, Rishi V. Puram, Francisca Vazquez, Barbara A. Weir, William C. Hahn, Benjamin Ebert & David Pellman. The EMT regulator ZEB2 is a novel dependency of human and murine acute myeloid leukemia. Blood 2017 Jan 26;129(4):497-508. doi: 10.1182/blood-2016-05-714493. Epub 2016 Oct 18.
Brenton R. Paolella, William J. Gibson, Laura M. Urbanski, John A. Alberta, Travis I. Zack, Pratiti Bandopadhayay, Caitlin A. Nichols, Pankaj K. Agarwalla, Meredith S. Brown, Rebecca Lamothe, Yong Yu, Peter S. Choi, Esther A. Obeng, Dirk Heckl, Guo Wei, Belinda Wang, Aviad Tsherniak, Francisca Vazquez, Barbara A. Weir, David E. Root, Glenn S. Cowley, Sara J. Buhrlage, Charles D. Stiles, Benjamin L. Ebert, William C. Hahn, Robin Reed, & Rameen Beroukhim. Copy-number and gene dependency analysis reveals partial copy loss of wild-type SF3B1 as a novel cancer vulnerability. Elife 2017 Feb 8;6. pii: e23268. doi: 10.7554/eLife.23268.
Jong Wook Kim, Olga B. Botvinnik, Omar Abudayyeh, Chet Birger, Joseph Rosenbluh, Yashaswi Shrestha, Mohamed E. Abazeed, Peter S. Hammerman, Daniel DiCara, David J. Konieczkowski, Cory M. Johannessen, Arthur Liberzon, Amir Reza Alizad-Rahvar, Gabriela Alexe, Andrew Aguirre, Mahmoud Ghandi, Heidi Greulich, Francisca Vazquez, Barbara A. Weir, Eliezer M. Van Allen, Aviad Tsherniak, Diane D. Shao, Travis I. Zack, Michael Noble, Gad Getz, Rameen Beroukhim, Levi A. Garraway, Masoud Ardakani, Chiara Romualdi, Gabriele Sales, David A. Barbie, Jesse S. Boehm, William C. Hahn, Jill P. Mesirov, & Pablo Tamayo. Characterizing genomic alterations in cancer by complementary functional associations. Nature Biotechnology 2016 May;34(5):539-46. doi: 10.1038/nbt.3527. Epub 2016 Apr 18.
Gregory V. Kryukov, Frederick H. Wilson, Jason R. Ruth, Joshiawa Paulk, Aviad Tsherniak, Sara E. Marlow, Francisca Vazquez, Barbara A. Weir, Mark E. Fitzgerald, Minoru Tanaka, Craig M. Bielski, Justin M. Scott, Courtney Dennis, Glenn S. Cowley, Jesse S. Boehm, David E. Root, Todd R. Golub, Clary B. Clish, James E. Bradner, William C. Hahn, & Levi A. Garraway. MTAP deletion confers enhanced dependency on the PRMT5 arginine methyltransferase in cancer cells. Science 2016 Mar 11;351(6278):1214-8. doi: 10.1126/science.aad5214. Epub 2016 Feb 11.
Hugh S. Gannon, Nathan Kaplan, Aviad Tsherniak, Francisca Vazquez, Barbara A. Weir, William C. Hahn & Matthew Meyerson. Identification of an "Exceptional Responder" Cell Line to MEK1 Inhibition: Clinical Implications for MEK-Targeted Therapy. Molecular Cancer Research 2016 Feb;14(2):207-15. doi: 10.1158/1541-7786.MCR-15-0321. Epub 2015 Nov 18. PMCID: PMC4755909.
Kimberly H. Kim, Woojin Kim, Thomas P. Howard, Francisca Vazquez, Aviad Tsherniak, Jennifer N. Wu, Weishan Wang, Jeffrey R. Haswell, Loren D. Walensky, William C. Hahn, Stuart H. Orkin, & Charles W. M. Roberts.SWI/SNF-mutant cancers depend on catalytic and non-catalytic activity of EZH2. Nature Medicine 2015 Dec;21(12):1491-6. doi: 10.1038/nm.3968. Epub 2015 Nov 9.
Mark M. Pomerantz, Fugen Li, David Y. Takeda, Romina Lenci, Apurva Chonkar, Matthew Chabot, Paloma Cejas, Francisca Vazquez, Jennifer Cook, Ramesh A. Shivdasani, Michaela Bowden, Rosina Lis, William C Hahn, Philip W. Kantoff, Myles Brown, Massimo Loda, Henry W. Long, & Matthew L. Freedman. The androgen receptor cistrome is extensively reprogrammed in human prostate tumorigenesis. Nature Genetics 2015 Nov;47(11):1346-51. doi: 10.1038/ng.3419. Epub 2015 Oct 12. PMCID: PMC4707683.
